The group investigates how different regulatory mechanisms affect information processing in developmental systems. We pursue that goal by exploring specific data-driven as well as general idealized models of cell fate specification. In the data-driven theme, we constrain models with high-resolution spatiotemporal data of spinal cord formation obtained from experimental labs. In the theory based theme, we construct models by combining general principles of morphogen production and spreading, with various mechanisms of interpretation of morphogen signalling by cells undergoing specification. By using these two complementary approaches we aim to understand the design principles of regulatory mechanisms that result in precise and reproducible pattern formation in wide range of developmental systems.
We also develop computational tools that facilitate exploration of multidimensional space of regulatory networks or genotypes to find solutions with the highest fitness against imposed functional goals.
Regulatory mechanisms for reproducible neural pattern specification in the developing spinal cord
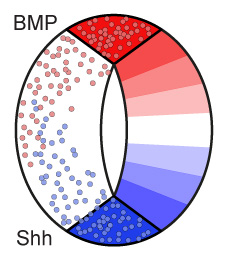
In the newly forming spinal cord, a striped pattern of distinct neural progenitor types is established. Despite the intrinsic stochasticity of signalling events and gene expression involved in pattern formation, the resulting spatial arrangement of neural progenitor types is remarkably precise and reproducible between individuals of the same species. How in the newly forming spinal cord the formation of the source region that secrets signalling molecules (morphogens), the spreading of morphogens, and signal interpretation by downstream target genes is coordinated is unclear. Our research goal is to identify emergent regulatory mechanisms involving signalling, gene regulation and growth that through a feedback loop between morphogen and target tissue result in precise pattern formation.
Information decoding in developmental systems
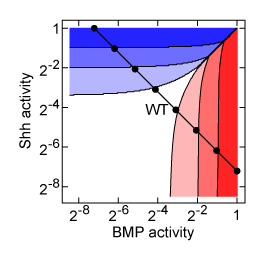
The development of organisms is a dynamic process in which cells proliferate, rearrange, and interpret molecular signals to adopt specific cell fates. The signals, so called morphogens, are interpreted to establish cell fate with striking precision of one cell diameter in Drosophila or three cell diameters in vertebrate spinal cord. How cells achieve this level of positional precision given the intrinsic stochasticity of signalling events and gene expression is still poorly understood. We approach this question by analysing how much positional information is available in the graded profiles of morphogen activity and how different modes of cell fate pattern specification process the available information. Our research goal is to identify regulatory mechanisms or modes of pattern specification that affect positional information decoding strategy in developmental systems.
Biophysical and functional constraints shape structure of regulatory networks
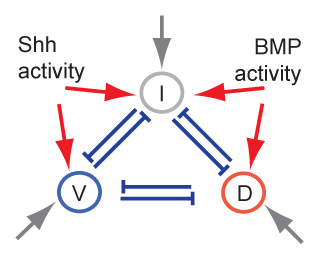
Gene regulatory networks (GRNs) allow to control the patterning of living cells. However, it is still unclear how specific circuitries of GRNs emerged in different biological systems. We have shown that small GRNs under mutation-selection balance with imposed biophysical and functional constraints can evolve towards their biological counterparts. In the case mimicking cell specification, the imposed condition of multiple stable gene expression patterns resulted in the overabundance of network motifs with mutual inhibitory interactions. In the more data-driven scenario of 3-gene network informed with spatiotemporal morphogen signalling the resulting GRN recapitulated the key experimental observations. Our research goal is to identify which biophysical and functional constraints are key determinants of GRNs observed in biological systems.
Evolutionary accessibility of highly fit regulatory mechanisms
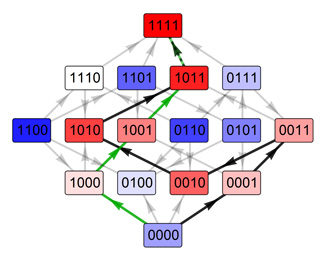
The genetic information describing genes and their regulatory interactions is encoded in the genotype. In the widely used case with binary genotypes (gene is either expressed or not) and randomly assigned fitness the best-adapted organism is almost always evolutionary inaccessible. Interestingly, if the number of units encoding genotype is increased to four DNA nucleotides (A,T,C,G) or ~20 amino acids used for synthesizing proteins, the probability for evolutionary process to find a way from lower-fitness organism to the best adapted organism is increased to more than 50%. However, this increase in accessibility comes with a price that evolutionary pathways become much longer and many mutations do not decrease a Hamming distance to the best adapted genotype. Our research goal is to understand these trade-off and other fitness landscape properties that render these landscapes easier to traverse by evolutionary process.
